11. Navigation and Layout¶
11.1. Basic Network Navigation¶
Cytoscape uses a Zoomable User Interface for navigating and viewing networks. ZUIs use two mechanisms for navigation: zooming and panning. Zooming increases or decreases the magnification of a view based on how much or how little a user wants to see. Panning allows users to move the focus of a screen to different parts of a view.
Zoom¶
Cytoscape provides four mechanisms for zooming: toolbar buttons, menu options, keyboard shortcuts and the scroll wheel.
Use the zooming buttons located on the toolbar to zoom in and out of the interaction network shown in the current network display. Zoom icons are detailed below:
![]()
From Left to Right:
Zoom In
Menu option: View → Zoom In
Keyboard shortcut:
Ctrl-Plus(Command-Pluson Mac OS X)
Zoom Out
Menu option: View → Zoom Out
Keyboard shortcut:
Ctrl-Minus(Command-Minuson Mac OS X)
Fit Content: Zoom level is adjusted to fit the network to the window
Menu option: View → Fit Content
Keyboard shortcut:
Ctrl-0(Command-0on Mac OS X)
Fit Selected: Zoom level is adjusted to zoom on any selected nodes
Menu option: View → Fit Selected
Keyboard shortcut:
Ctrl-9(Command-9on Mac OS X)
Using the scroll wheel, you can zoom in by scrolling up and zoom out by scrolling downwards. These directions are reversed on Macs with natural scrolling enabled (the default for Mac OS X Lion and newer versions).
Pan¶
There are two ways to pan the network:
Left-Click and Drag - You can pan the network view by holding down the left mouse button and moving the mouse.
Dragging Box on Network Overview - You can also pan the view by left-clicking and dragging the blue box in the overview panel in the lower part of the view.
11.2. Other Mouse Behaviors¶
Select¶
Click the left mouse button on a node, edge or annotation to select that element.
Hold down the
ShiftorCtrlkey (Commandon Mac) and left-click a node, edge or annotation to add it to the selection. Doing the same on a selected element unselects it.Hold down the left mouse button on the canvas background and drag the mouse while holding down the
ShiftorCtrlkey (Commandon Mac) to select groups of nodes/edges/annotations using a classic rectangular selection area.Hold down the left mouse button on the canvas background and drag the mouse while holding down
Option+Shift(Command+Shifton Mac) to select groups of nodes/edges/annotations using a lasso selection tool.Remember that the selection action (mouse click or drag-selection) only works if the Selection Mode for that element type (i.e. nodes, edges, annotations) is enabled. In order to enable or disable the selection of an element type, just toggle its corresponding button at the bottom of the network view (see image below) or use the options under the menu Select → Mouse Drag Selects.
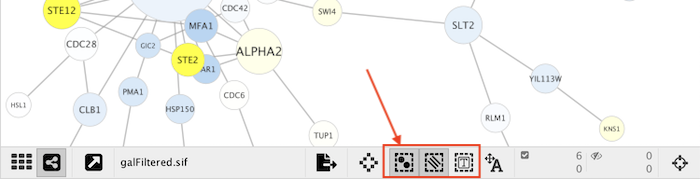
So if you don’t want any nodes to be selected, toggle the button
 off.
off.And if you don’t want any edges to be selected, toggle the button
 off.
off.Likewise, if you don’t want any annotations to be selected, toggle the button
 off.
off.
Context¶
Click the right mouse button (or Ctrl+left mouse button on Macs) on a node/edge to launch a context-sensitive menu with additional information about the node/edge.
Node Context Menu¶
This menu can change based on the current context. For nodes, it typically shows:
Add
Diffuse
Edit
Select
Group
Nested Networks
Apps
External Links
Preferences
Edges usually have the following menu:
Diffuse
Edit
Select
Apps
External Links
Preferences
Apps can contribute their own items into node and edge context menus. These additions usually appear in the Apps section of the context menu.
Nested Network Node Context Menu¶
Add Nested Network: Lets the user select any network in Cytoscape as the current node’s nested network. If the current node already has a nested network it will be replaced.
Remove Nested Network: Removes the currently associated nested network from a node. The associated network is not deleted. Only the association between the node and the network is removed.
Go to Nested Network: The current node’s nested network will be the current network view and have the focus. Should a network view for the nested network not exist, it will be created.
More information about nested networks can be found in the Nested Networks section.
11.3. Manual Layout¶
The simplest method to manually organize a network is to click on a node and drag it. All of the selected nodes are moved together.
In addition to the ability to click on a node and drag it to a new position, Cytoscape now has the ability to move nodes using the arrow keys on the keyboard. By selecting one or more nodes using the mouse and clicking one of the arrow keys (←, ↑, →, ↓) the selected nodes will move one pixel in the chosen direction. If an arrow key is pressed while holding the Shift key down, the selected nodes will move 15 pixels in the chosen direction.
11.4. Node Layout Tools¶
The Tool Panel is available via the menu command View → Show Tool Panel, or via Layout → Node Layout Tools.
It contains several Node Layout Tools that can help to automate or fine tune a layout.
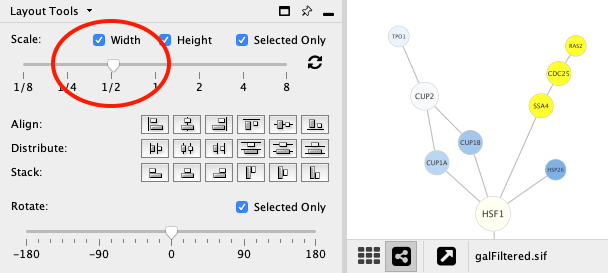
Scale¶
Adjust the Scale slider to change the length of edges. The position of the nodes will be scaled, not the node sizes. Node size can be adjusted using Styles. The images below show selected (yellow) nodes scaled to 50% of the default value.
Before
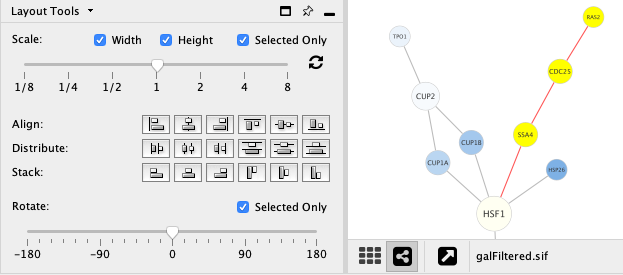
After
Rotate¶
The Rotate function will change the orientation of the entire network or a selected portion of the network. The images below show a network with selected nodes rotated 90 degrees.
Before

After
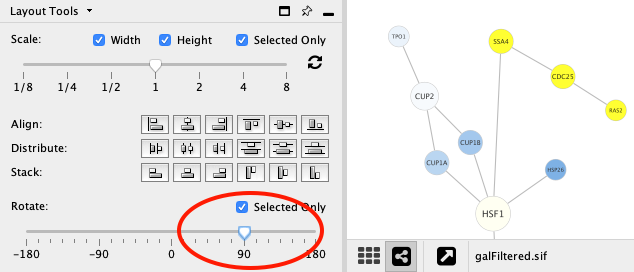
Align, Distribute and Stack¶
The Tool Panel contains buttons to set relative postions of nodes.
Align provides different options for either vertically or horizontally aligning selected nodes against a line. The differences are in what part of the node gets aligned, e.g. the center of the node, the top of the node, the left side of the node. Distribute evenly distributes selected nodes between the two most distant nodes along either the vertical or horizontal axis. The differences are again a function what part of the node is used as a reference point for the distribution. Stack vertically or horizontally stacks selected nodes with the full complement of alignment options. The table below provides a description of what each button does.
| Button | Before | After | Description of Align Options |
|---|---|---|---|
 |  |  | Vertical Align Top - The tops of the selected nodes are aligned with the top-most node. |
 |  |  | Vertical Align Center - The centers of the selected nodes are aligned along a line defined by the midpoint between the top and bottom-most nodes. |
 |  |  | Vertical Align Bottom - The bottoms of the selected nodes are aligned with the bottom-most node. |
 |  |  | Horizontal Align Left - The left hand sides of the selected nodes are aligned with the left-most node. |
 |  |  | Horizontal Align Center - The centers of the selected nodes are aligned along a line defined by the midpoint between the left and right-most nodes. |
 |  |  | Horizontal Align Right - The right hand sides of the selected nodes are aligned with the right-most node. |
| Button | Before | After | Description of Align Options |
|---|---|---|---|
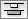 |  | 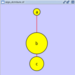 | Vertical Distribute Top - The tops of the selected nodes are distributed evenly between the top-most and bottom-most nodes, which should stay stationary. |
 |  |  | Vertical Distribute Center - The centers of the selected nodes are distributed evenly between the top-most and bottom-most nodes, which should stay stationary. |
 |  | 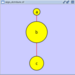 | Vertical Distribute Bottom - The bottoms of the selected nodes are distributed evenly between the top-most and bottom-most nodes, which should stay stationary. |
 | 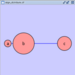 | 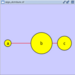 | Horizontal Distribute Left - The left hand sides of the selected nodes are distributed evenly between the left-most and right-most nodes, which should stay stationary. |
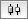 |  | 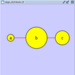 | Horizontal Distribute Center - The centers of the selected nodes are distributed evenly between the left-most and right-most nodes, which should stay stationary. |
 |  | 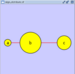 | Horizontal Distribute Right - The right hand sides of the selected nodes are distributed evenly between the left-most and right-most nodes, which should stay stationary. |
| Button | Before | After | Description of Align Options |
|---|---|---|---|
 | 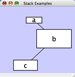 | 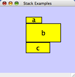 | Vertical Stack Left - Vertically stacked below top-most node with the left-hand sides of the selected nodes aligned. |
 |  | 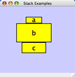 | Vertical Stack Center - Vertically stacked below top-most node with the centers of selected nodes aligned. |
 |  | 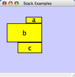 | Vertical Stack Right - Vertically stacked below top-most node with the right-hand sides of the selected nodes aligned. |
 | 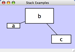 | 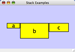 | Horizontal Stack Top - Horizontally stacked to the right of the left-most node with the tops of the selected nodes aligned. |
 |  | 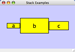 | Horizontal Stack Center - Horizontally stacked to the right of the left-most node with the centers of selected nodes aligned. |
 |  | 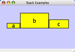 | Horizontal Stack Bottom - Horizontal Stack Center - Horizontally stacked to the right of the left-most node with the bottoms of the selected nodes aligned. |
11.5. Edge Bend and Automatic Edge Bundling¶
From Cytoscape 3.0, Edge Bend is a regular edge property and can be used as a part of a Style. Just like any other edge property, you can select a Default Value, a Mapping and use Bypass for select nodes. In the Style tab, select the Bend property from the Properties drop-down and click on either the Default Value, Mapping or Bypass cell to bring up the Edge Bend Editor. In the editor, you can add as many handles as you want to the edge using Alt-Click on Windows, Option-Click on Mac, or Ctrl-Alt-Click on Linux.
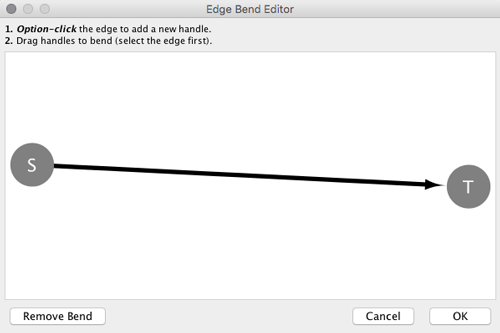
To clear all edge bends, select Layout → Clear All Edge Bends.
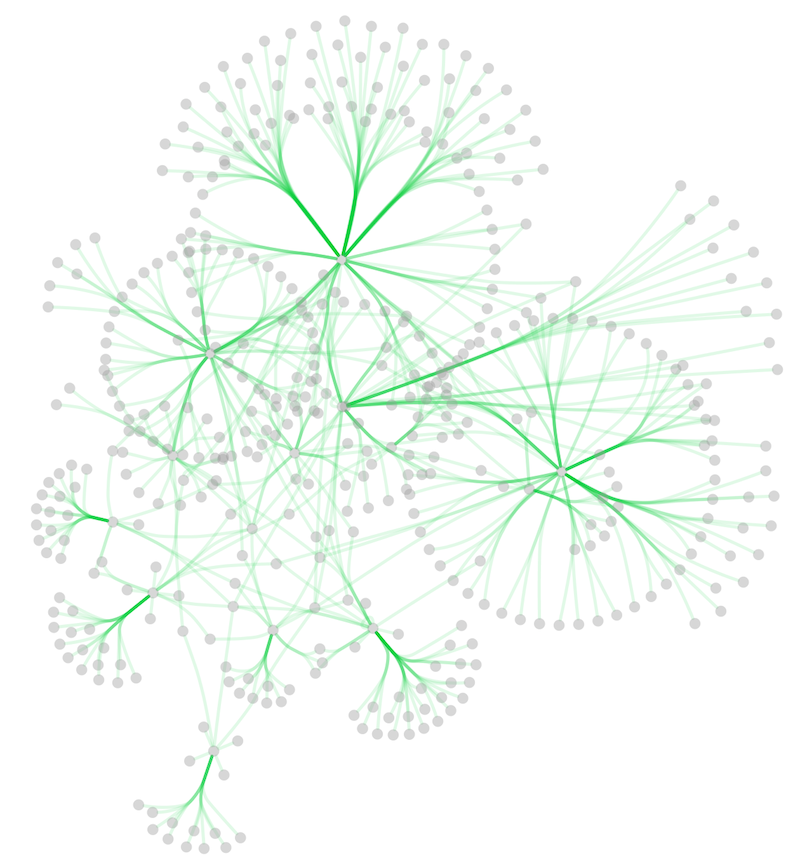
In addition to adding handles manually, you can use the Bundle Edges function to bundle all or selected edges automatically.
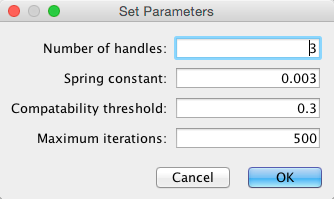
Select Layout → Bundle Edges → All Nodes and Edges.
Set parameters.
Details of the algorithm is described in this paper.
Press OK to run. Edge bundling may take a long time if the number of edges is large.
If it takes too long, try decreasing Maximum Iterations.
For large, dense networks, try setting Maximum iterations in the range of 500 - 1000.
Note: The handle locations will be optimized for current location of nodes. If you move node positions, you need to run the function again to get proper result.
11.6. Automatic Layout Algorithms¶
The Layout menu has an array of features for organizing the network visually according to one of several algorithms, aligning and rotating groups of nodes, and adjusting the size of the network. Cytoscape layouts have three different sources, which are reflected in the Layout menu.
Cytoscape layouts have the option to operate on only the selected nodes, and all provide a Settings… panel to change the parameters of the algorithm. Most of the Cytoscape layouts also partition the graph before performing the layout. In addition, many of these layouts include the option to take either node or edge columns into account. A few of the layout algorithms are:
Grid Layout¶
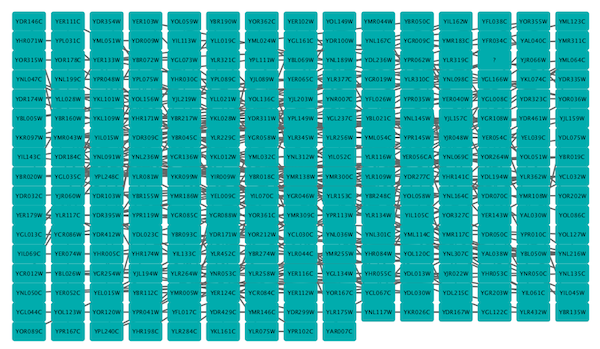

The grid layout is a simple layout the arranges all of the nodes in a square grid. This is the default layout and is always available as part of the Cytoscape core. It is available by selecting Layout → Grid Layout. A sample screen shot is shown above.
Edge-weighted Spring-Embedded Layout¶
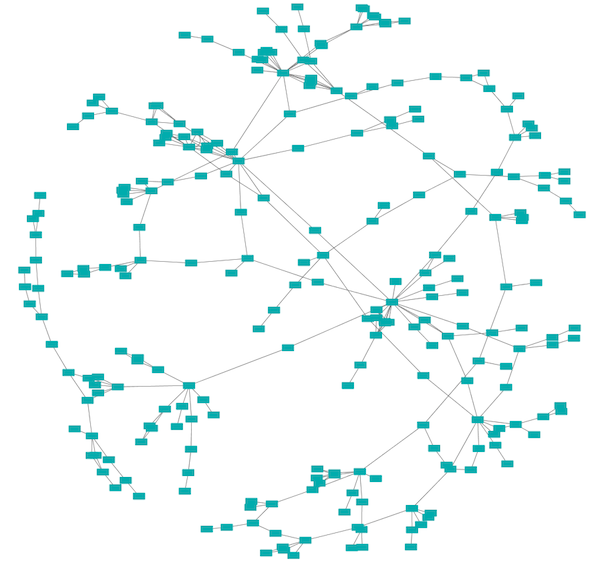
The spring-embedded layout is based on a force-directed paradigm as implemented by Kamada and Kawai (1988). Network nodes are treated like physical objects that repel each other, such as electrons. The connections between nodes are treated like metal springs attached to the pair of nodes. These springs repel or attract their end points according to a force function. The layout algorithm sets the positions of the nodes in a way that minimizes the sum of forces in the network. This algorithm can be applied to the entire network or a portion of it by selecting the appropriate options from Layout → Edge-weighted Spring Embedded.
Attribute Circle Layout¶
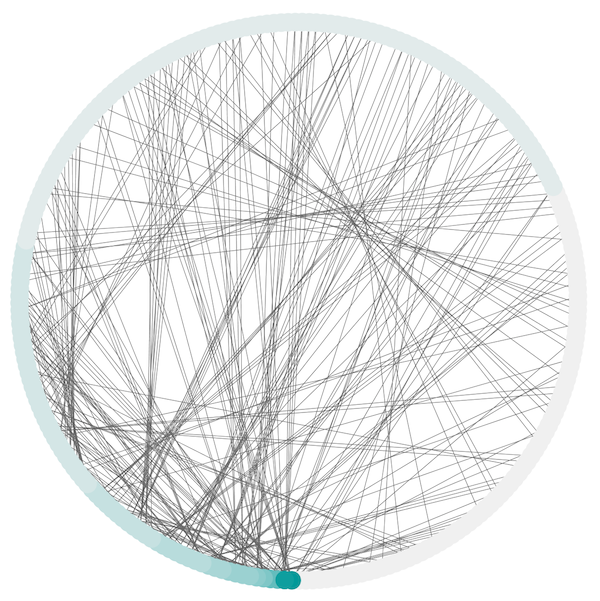

The Attribute Circle layout is a quick, useful layout, particularly for small networks, that will locate all of the nodes in the network around a circle. The node order is determined by a user-selected node column. The result is that all nodes with the same value for that column are located together around the circle. Using Layout → Attribute Circle Layout → column to put all nodes around a circle using column to position them. The sample screen shot above shows the a subset of the galFiltered network organized by node degree.
Group Attributes Layout¶
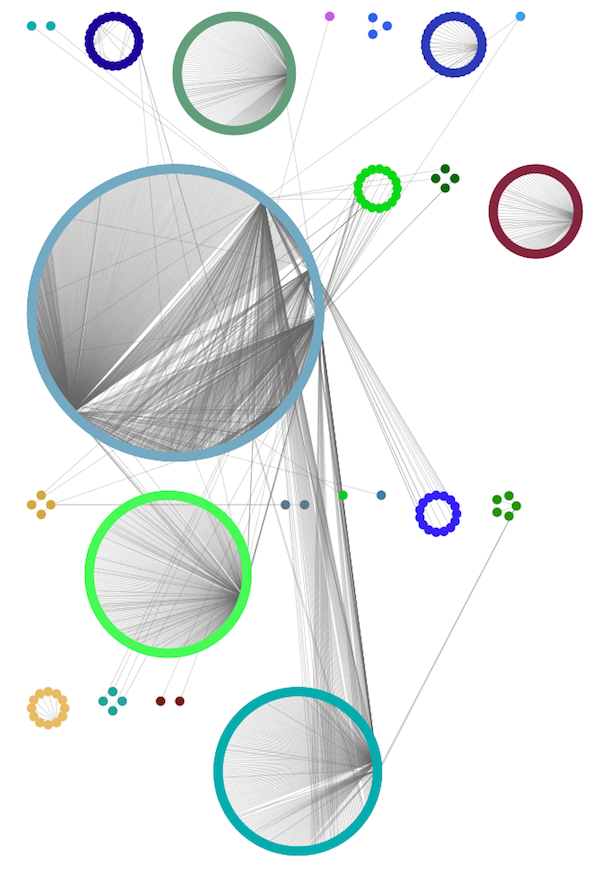

The Group Attributes layout is similar to the Attribute Circle layout described above except that instead of a single circle with all of the nodes, each set of nodes that share the same value for the column are laid out in a separate circle. The same network shown above (network generated by PSICQUIC Client) is shown above, using Layout → Group Attributes Layout → taxonomy.
Prefuse Force Directed Layout¶

The force-directed layout is a layout based on the force-directed paradigm. This layout is based on the algorithm implemented as part of the prefuse toolkit provided by Jeff Heer. The algorithm is very fast and with the right parameters can provide a very visually pleasing layout. The Force Directed Layout will also accept a numeric edge column to use as a weight for the length of the spring, although this will often require more use of the Settings… dialog to achieve the best layout. This algorithm is available by selecting Layout → Prefuse Force-Directed Layout → (unweighted) or the edge column you want to use as a weight. A sample screen shot showing a portion of the galFiltered network provided in sample data is provided above.
Compound Spring Embedder Layout¶
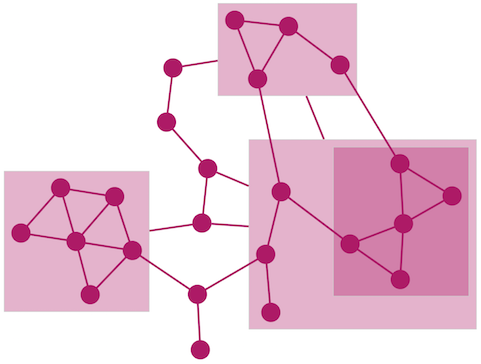
The Compound Spring Embedder (CoSE) layout is based on the traditional force-directed layout algorithm with extensions to handle multi-level nesting (compound nodes), edges between nodes of arbitrary nesting levels and varying node sizes. It is the suggested Cytoscape layout for compound graphs, although it also works very well with noncompound graphs. It is available by selecting Layout → Compound Spring Embedder (CoSE).
Circular Layout¶
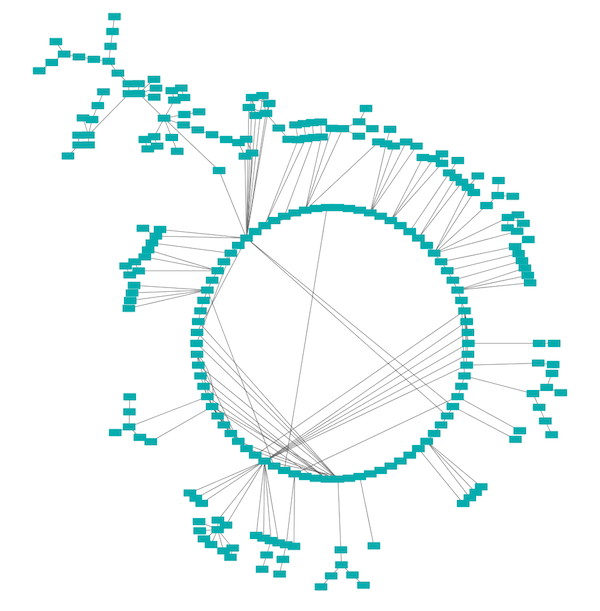

This algorithm produces layouts that emphasize group and tree structures within a network. It partitions the network by analyzing its connectivity structure, and arranges the partitions as separate circles. The circles themselves are arranged in a radial tree layout fashion. This algorithm is available by selecting Layout → Circular Layout.
Hierarchical Layout¶
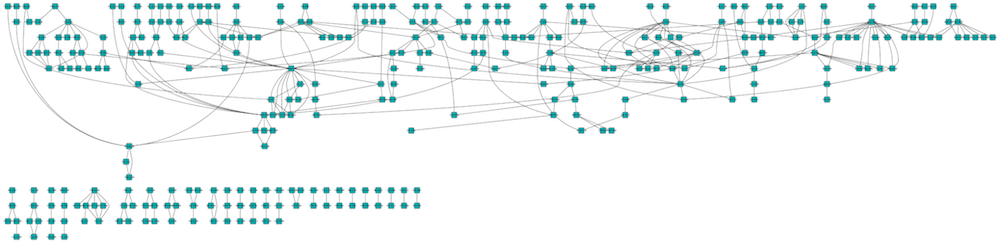
The hierarchical layout algorithm is good for representing main direction or flow within a network. Nodes are placed in hierarchically arranged layers and the ordering of the nodes within each layer is chosen in such a way that minimizes the number of edge crossings. This algorithm is available by selecting Layout → Hierarchical Layout.
Copycat Layout¶
The Copycat layout uses node positions in one network to lay out nodes in another network. Selecting Layout → Copycat Layout displays a dialog box that allows you to select the source network (which is already laid out) and the target network (which needs layout). By default, Copycat matches nodes in the source and target networks by node name, but you can choose any node attribute for this match.
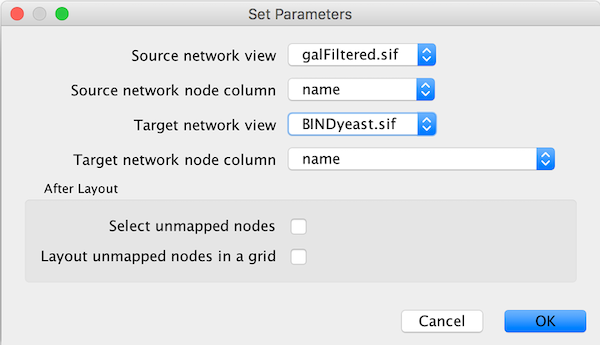
As options, you can use Select unmapped nodes to cause Copycat to select target nodes that were not matched by nodes in the source network. This allows you to move them and possibly lay them out using other layout algorithms. You can use the Layout unmapped nodes in a grid option to preemptively move the unmatched target nodes away from the laid out target network.
11.7. yFiles Layouts¶
Cytoscape offers a set of layout algorithms based on the yFiles library. As of Cytoscape 3.6, the yFiles algorithms are available directly through the Cytoscape App Store. To install them, go to Layout → Install yFiles, which will direct you to the App Store. Click the Install button to proceed. You will be directed to a license agreement, and once yFiles is installed, the yFiles layouts will be available in the Layout menu.
The layout algorithms included in yFiles are:
Circular Layout
Hierarchic Layout
Hierarchic Layout Selected Nodes
Organic Layout
Orthogonal Layout
Radial Layout
Tree Layout
Orthogonal Edge Router
Organic Edge Router
Below are a few examples of yFiles layouts in Cytoscape, with more available from yFiles.
yFiles Organic Layout¶
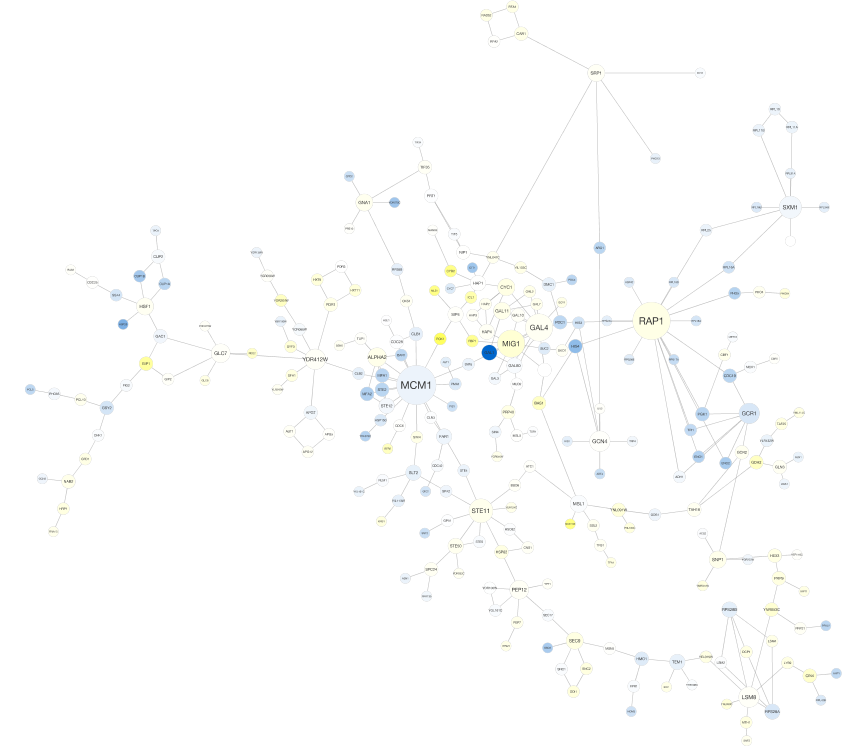
yFiles Hierarchic Layout¶
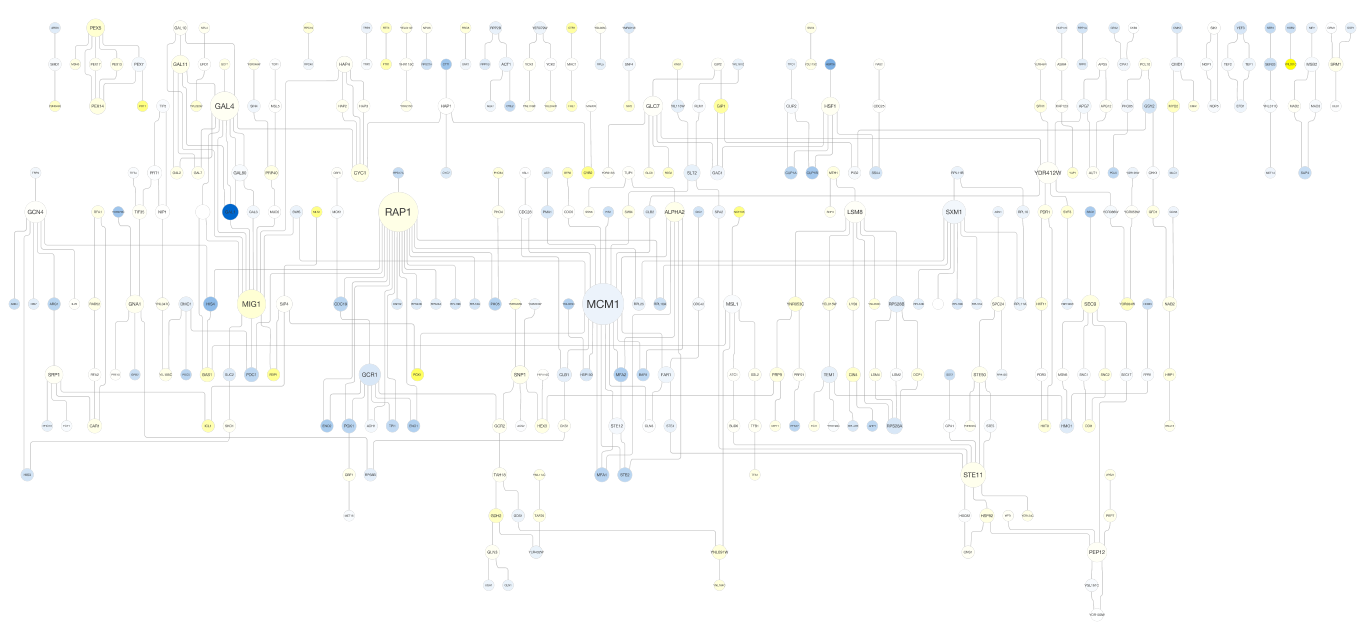
yFiles Tree Layout¶
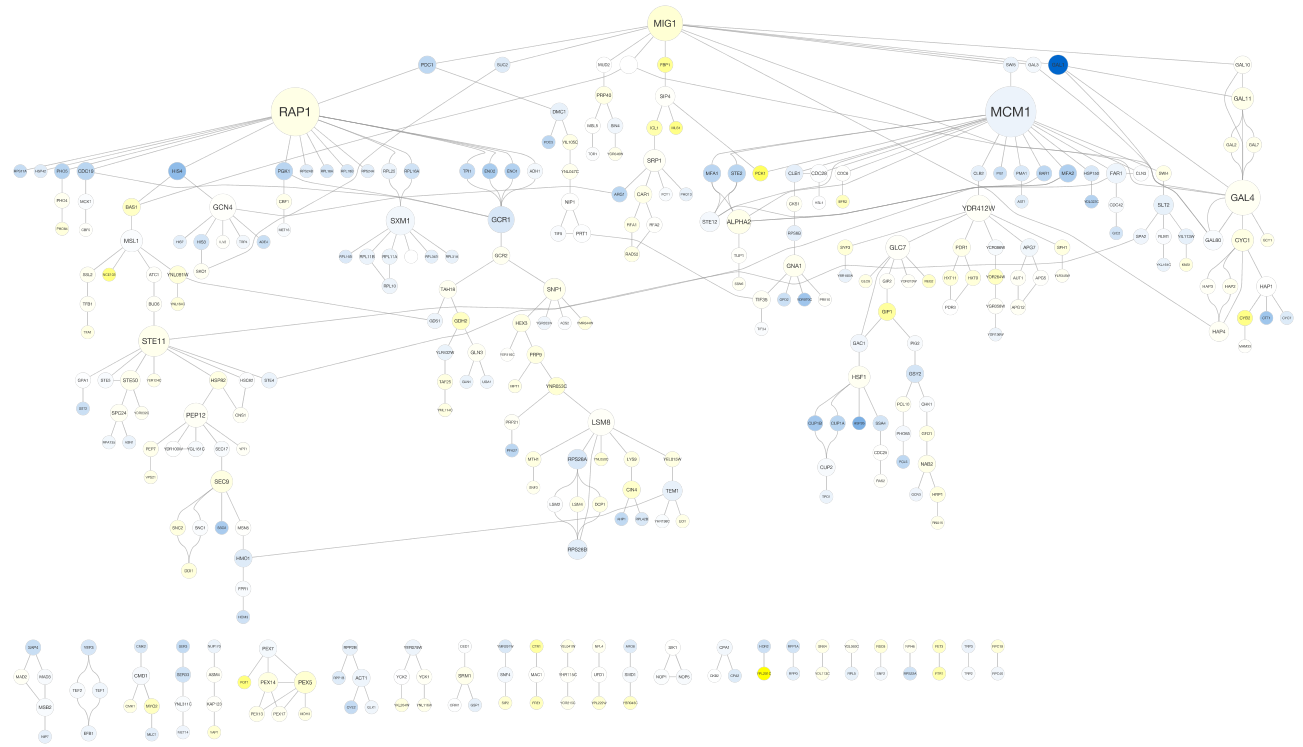
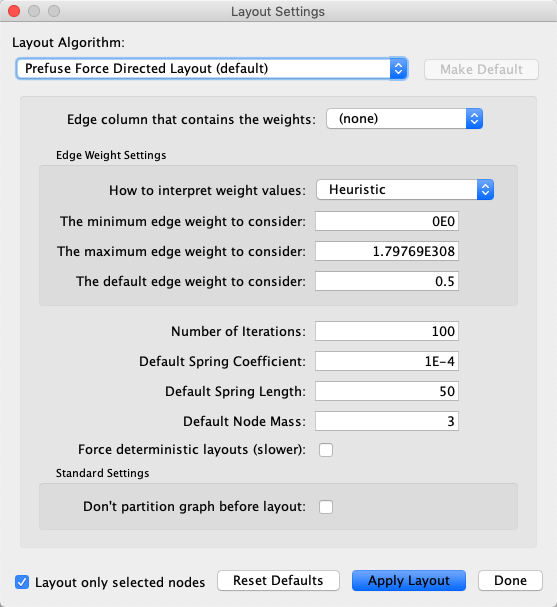
11.8. Layout Parameters¶
Many layouts have adjustable parameters that are exposed through the Layouts → Settings… menu option. The Layout Settings dialog, which allows you to choose which layout algorithm settings to adjust, is shown below. The settings presented vary by algorithm and only those algorithms that allow access to their parameters will appear in the drop-down menu at the top of the dialog. Once you’ve modified a parameter, clicking the Execute Layout button will apply the layout.