17. Analyzer¶
Analyzer computes a comprehensive set of topological metrics for undirected and directed networks, including:
Number of nodes, edges and connected components.
Network diameter, radius and clustering coefficient, as well as the characteristic path length.
Charts for topological coefficients, betweenness, and closeness.
Distributions of degrees, neighborhood connectiveness, average clustering coefficients, shortest path lengths, number of shared neighbors and stress centrality.
17.1. Network Analysis¶
Analyze Network¶
To run Analyzer, select Tools → Analyze Network.
Analyzer will run different statistics depending on whether the network is directed or undirected, with the default being undirected. A Set Parameters dialog will open where you can specify if the network should be analyzed as a directed graph.
When results are ready, they will appear in the Results Panel.
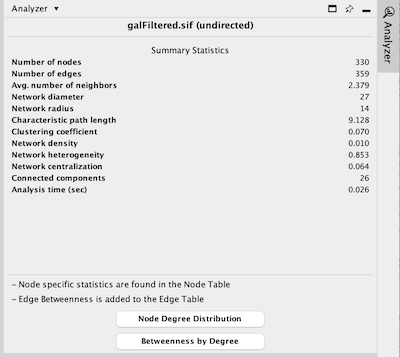
The results have multiple tabs, accessible via the buttons on the bottom of the panel. Details on the network metrics can be found here.
Simple Metrics
Node Degree Distribution
Clustering Coefficient
Topological Coefficient
Average Shortest Path Length
Neighborhood Connectivity
Betweenness Centrality
Closeness Centrality
Stress Centrality
Analyze Subset of Nodes¶
Prior versions of this tool offered the option of analyzing all nodes or only a selected subset. This is no longer supported directly in the program. Instead, if you want to analyze a subnetwork, you can use the command File → New Network → From Selected Nodes, All Edges in the Command Panel to create the desired subnetwork.
Plot Metrics¶
Once the Analyzer is run, several additional columns are added to the Node Table (and an EdgeBetweenness column is added to the Edge Table). To plot any of these new columns, right-click on the column header and select Plot Histogram… for a single parameter distribution, or Plot Scatter… for a bivariate plot of the data. Within either of these charts it is possible to select a section of the data, and select the nodes (edges) in the main graph window corresponding to the region selected on the chart.
For additional information about NetworkAnalyzer see Yassen Assenov, Nadezhda Doncheva, Thomas Lengauer, and Mario Albrecht, DOI: 10.1038/nprot.2012.004.